## Install R packages
# May need to install these operating system packages (if using Ubuntu)
# sudo apt-get install libabsl-dev cmake
# The following installs the latest stable R package versions from the CRAN repository (install.packages) or the latest development code from Github (remotes::install_github)
if (!require(remotes)) {
install.packages("remotes")
}
library(remotes)
if (!require(terra)) {
remotes::install_github("rspatial/terra")
}
if (!require(geodata)) {
install.packages("geodata")
}
if (!require(magick)) {
install.packages("magick")
}
# Load libraries
library(terra)
library(geodata)
library(magick)
# Directories to store WorldClim data
out_dir <- "wc_downloads_1_khm"
dir.create(out_dir, showWarnings = FALSE)
out_dir_historical <- "wc_downloads_1_khm/historical"
dir.create(out_dir_historical, showWarnings = FALSE)
out_dir_future <- "wc_downloads_1_khm/future"
dir.create(out_dir_future, showWarnings = FALSE)
# Directory to store maps and graphic images
out_dir_graphics <- "wc_downloads_1_khm/graphics"
dir.create(out_dir_graphics, showWarnings = FALSE)A Transferable Climate Profile Framework
Using WorldClim v2.1 and CMIP6 SSP Projections
Case Study: Cambodia
Introduction
A climate profile is an essential element of any climate risk assessment, as it provides the quantitative baseline and future projections necessary to identify climate hazards, evaluate exposure, and inform adaptation planning. This climate profile is part of a larger framework under development for assessing climate risk, which will be shared once completed.
This R script provides a transparent, reproducible and transferable framework for generating a set of key maps and graphs required for a climate profile. Outputs are based on the WorldClim 2.1 climate dataset, including historical (1970–2000) and CMIP6 projected data for two future periods (2041–2060 and 2061–2080) under two shared socioeconomic pathways (SSP): SSP2-4.5 (moderate) and SSP5-8.5 (very high) emission scenarios. Future projections for Cambodia are based on an ensemble of five global climate models (GCMs)—ACCESS-CM2, MIROC6, EC-Earth3-Veg, MRI-ESM2-0, and IPSL-CM6A-LR (see index.Rmd). While demonstrated here at the national level, the boundary used to crop data intputs can be readily adjusted to any administrative region or river basin boundary, making the script adaptable to sub-national and watershed-level climate assessments.
WorldClim is a popular open-source climate dataset, recognised for its global coverage and fine spatial resolution (1 km). However, as data are provided as monthly means, they cannot support derivation of daily-based climate change indices, such as heavy precipitation frequency, the number of consecutive dry days, or percentile-based metrics (e.g., as defined by the ETCCDI).1 Datasets such as ERA5 or CHIRPS with daily data increments could be used to generate daily-based climate change indices.
This climate profile presents historical and future projections of temperature and precipitation for Cambodia. The historical climate (1970–2000) is characterised through gridded spatial maps of annual mean temperature and precipitation, and a monthly precipitation bar chart illustrating the monsoon pattern with a distinct wet season (May–October) peaking in September and October and a dry season (November–April) (RIMES & UNDP, 2020). Future projections under SSP2-4.5 and SSP5-8.5 for 2041–2060 and 2061–2080 are presented through four complementary visualisations: spatial anomaly panel maps of annual mean temperature and precipitation changes; ensemble range plots of monthly mean temperature and precipitation with GCM uncertainty envelopes; and model ensemble spread plots of annual mean temperature and precipitation showing projected anomalies and model uncertainty across the five GCMs.
The climate profile outputs are categorised into three sections, including monthly and annual maps and graphics that can be used anywhere in the world, as well as a third section that is tailored to the region and sector of interest, in this case Cambodia and rice farming.
Prerequisites
Install and load R packages.
Download GCM data: five models, two scenarios and two future periods
The selection of models used is based on a preliminary review of literature pertinent to the case study area of Cambodia.
# Modify with country ISO code (used to download country boundary)
country_iso <- "KHM"
# Specify lat and lon of area of WorldClim data to be downloaded
lat <- 8
lon <- 100
gcms <-
c(
"ACCESS-CM2",
"MIROC6",
"EC-Earth3-Veg",
"MRI-ESM2-0",
"IPSL-CM6A-LR"
)
ssps <- c("245", "585")
years <- c("2041-2060", "2061-2080")
vars <- c("tmin", "tmax", "prec")
# Read in national boundary of interest
adm0 <- gadm(
country = country_iso,
level = 0,
path = out_dir
)Download historical data
Historical climate data for WorldClim covers the period from 1970-2000. Spatial resolution downloaded is 0.00833\(\dot{3}\)° (~1 km at equator).
for (v in vars) {
# Check if already downloaded
existing <-
list.files(
out_dir_historical,
pattern = paste0("wc2\\.1_30s_", v, ".*\\.tif$"),
full.names = TRUE
)
if (length(existing) > 0) {
message("Skipping ", v, " - already downloaded: ", basename(existing[1]))
next
}
message("Downloading ", v, " historical")
tmpdir <- tempfile(pattern = "wcdl_")
dir.create(tmpdir)
worldclim_tile(
var = v,
lon = lon,
lat = lat,
path = tmpdir
)
tifs <-
list.files(tmpdir,
pattern = "\\.tif$",
full.names = TRUE,
recursive = TRUE
)
if (length(tifs) == 0) {
stop("No tif downloaded for ", v)
}
src <- tifs[1]
newname <-
file.path(out_dir_historical, paste0(basename(src)))
result <- file.copy(src, newname, overwrite = TRUE)
if (result == FALSE) {
stop("Copy failed for ", src)
}
unlink(tmpdir, recursive = TRUE)
}Download future projections
Spatial resolution downloaded is 0.00833\(\dot{3}\)° (~1 km at equator).
# Download and store GeoTIFFs (per GCM, variable, SSP, and period)
for (g in gcms) {
for (s in ssps) {
for (v in vars) {
for (y in years) {
# Check if already downloaded
existing <- list.files(out_dir_future, pattern = paste0("wc2\\.1_30s_", v, "_", g, "_", "ssp", s, "_", y, ".*\\.tif$"), full.names = TRUE)
if (length(existing) > 0) {
message("Skipping ", v, " ", g, " ", s, " ", y, " - already downloaded")
next
}
message("Downloading ", v, " for GCM ", g)
tmpdir <- tempfile(pattern = "wcdl_")
dir.create(tmpdir)
geodata::cmip6_tile(
lon = lon, lat = lat, res = 0.5, path = tmpdir,
model = g, ssp = s, var = v, time = y
)
tifs <- list.files(tmpdir, pattern = "\\.tif$", full.names = TRUE, recursive = TRUE)
if (length(tifs) == 0) stop("No tif downloaded for ", v, " ", g)
src <- tifs[1]
newname <- file.path(out_dir_future, paste0(basename(src)))
result <- file.copy(src, newname, overwrite = TRUE)
if (result == FALSE) stop("Copy failed for ", src)
unlink(tmpdir, recursive = TRUE)
}
}
}
}Annual maps and graphics
The following can be used to generate annual maps and graphics for any region of the world.
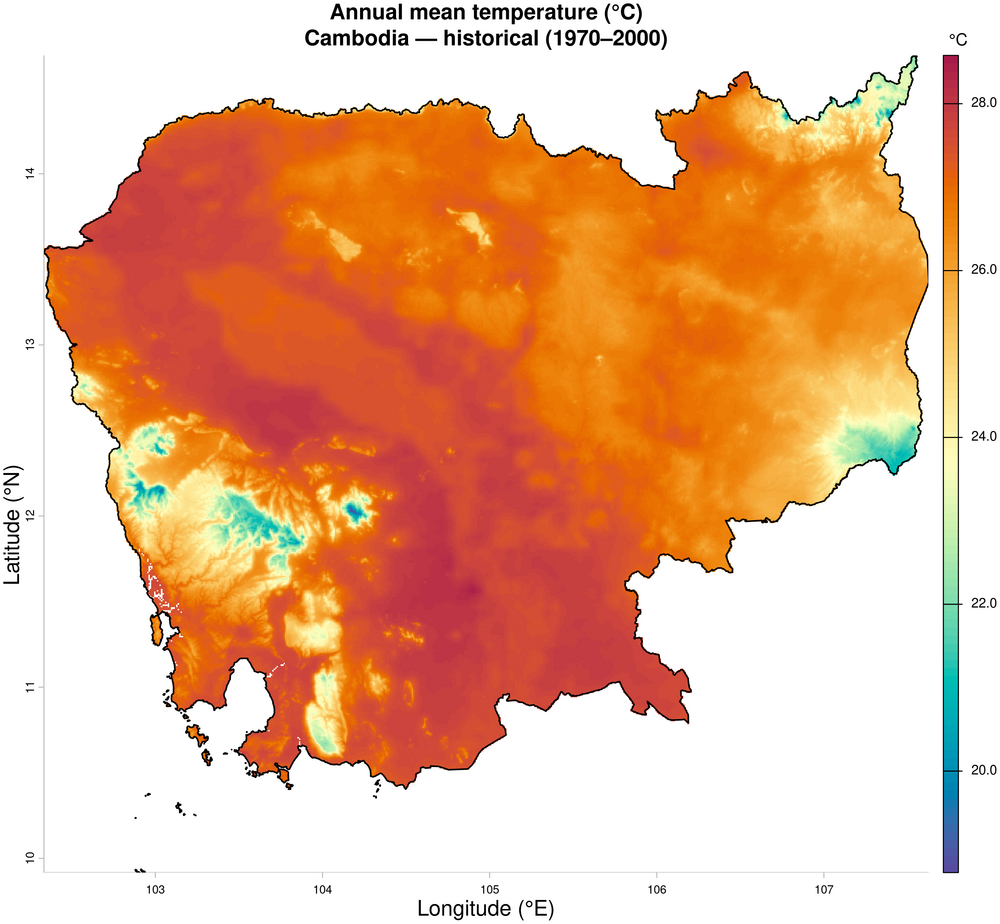
1. Annual mean temperature — historical baseline (1970–2000)
Process:
- Read in tmax and tmin GeoTIFFs (12 monthly layers in each GeoTIFF)
- Calculate annual mean temperatures from tmin and tmax monthly values: output two layers
- Calculate mean annual temperature from annual tmin and tmax means
# ============================================================
# Annual mean temperature — historical (1970–2000)
# Output: Gridded annual mean temperature map
# ============================================================
# Read in historical data
hist_tmin <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_tmin.tif"))
hist_tmax <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_tmax.tif"))
# Crop to Cambodia boundary
adm0 <- geodata::gadm(country = "KHM", level = 0, path = out_dir)
hist_tmin_c <- crop(hist_tmin, adm0) |> mask(adm0)
hist_tmax_c <- crop(hist_tmax, adm0) |> mask(adm0)
# Calculate mean temperature
tmin_annual <- app(hist_tmin_c, mean)
tmax_annual <- app(hist_tmax_c, mean)
tmean_annual <- (tmin_annual + tmax_annual) / 2
## Plot
# Save output as PNG
fname <- paste0(out_dir_graphics, "/annual_mean_temp_historical")
png(paste0(fname, ".png"), width = 12, height = 10, units = "in", res = 300, type = "cairo")
# To output PDF instead of PS/EPS outputs
# Comment out the cairo_ps line and uncomment the cairo_pdf lines below
# cairo_ps(paste0(out_dir_graphics, "/cambodia_tmean_historical.ps"), width = 8, height = 7)
# cairo_pdf(paste0(out_dir_graphics, "/cambodia_tmean_historical.pdf"), width = 8, height = 7)
plot(tmean_annual,
main = "Annual mean temperature (°C)\nCambodia — historical (1970–2000)",
col = hcl.colors(100, "Spectral", rev = TRUE),
mar = c(5.1, 4.1, 4.1, 2.1),
xlab = "Longitude (°E)",
ylab = "Latitude (°N)",
cex.main = 1.3,
cex.lab = 1.3,
plg = list(title = "°C"),
box = FALSE
)
lines(adm0, col = "black", lwd = 1.5)
dev.off()
# Convert PS to EPS file (use if inserting into R Markdown file
# to automatically remove white space)
# Comment these lines out if PDF output is specified above
# if (!file.exists(paste0(fname, ".eps"))) {
# system(paste0("ps2eps -f ", fname, ".ps"))
# }
2. Annual mean precipitation — historical baseline (1970–2000)
Process:
- Read in precipitation GeoTIFF (12 monthly layers)
- Calculate annual mean precipitation (sum months)
# ============================================================
# Annual mean precipitation — historical (1970–2000)
# Output: Gridded annual mean precipitation map
# ============================================================
# Read in historical data
hist_prec <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_prec.tif"))
# Crop to Cambodia boundary
adm0 <- geodata::gadm(country = "KHM", level = 0, path = out_dir)
hist_prec_c <- crop(hist_prec, adm0) |> mask(adm0)
# Total annual mean precipitation
# Sum across 12 months for total annual precipitation
prec_annual <- app(hist_prec_c, sum)
## Plot
# Save output as PNG
fname <- paste0(out_dir_graphics, "/annual_mean_prec_historical")
png(paste0(fname, ".png"), width = 12, height = 10, units = "in", res = 300, type = "cairo")
plot(prec_annual,
main = "Annual mean precipitation (mm)\nCambodia \u2014 historical (1970\u20132000)",
col = hcl.colors(100, "YlGnBu", rev = TRUE),
mar = c(5.1, 4.1, 4.1, 2.1),
xlab = "Longitude (\u00B0E)",
ylab = "Latitude (\u00B0N)",
cex.main = 1.3,
cex.lab = 1.3,
plg = list(title = "mm"),
box = FALSE
)
lines(adm0, col = "black", lwd = 1.5)
dev.off()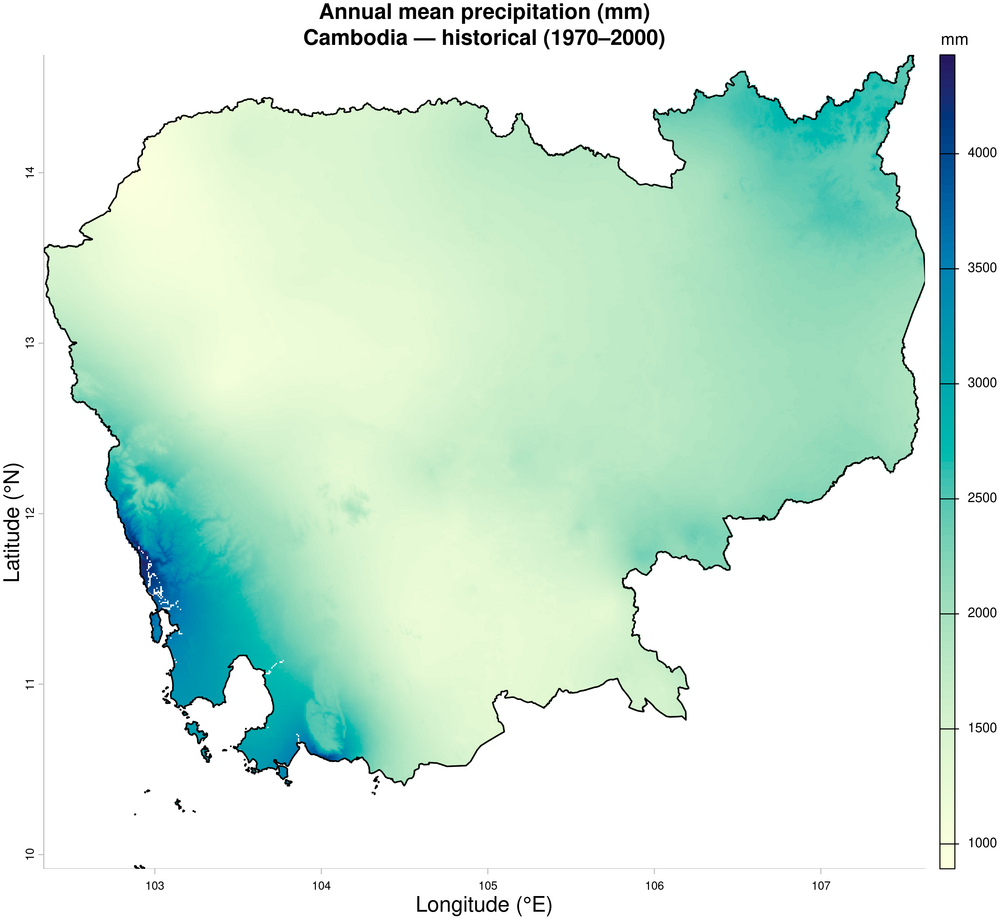
3. Annual mean temperature change — future (ensemble mean) minus historical
Temperature change maps are produced with a shared colour scale for cross-panel comparison.
Process: 1. Read in tmin and tmax for each of the 5 GCMs 2. Calculate temperature mean for each GCM: output 5 separate layers 3. Stack the 5 layers with rast(gcm_layers) 4. Take the mean across the 5 GCMs with app(…, mean): output single layer 5. Subtract historical annual mean: output anomaly map
# ============================================================
# Annual mean temperature change — 2x2 grid (SSP x Year)
# Output: Spatial anomaly panel map
# ============================================================
# Calculate all anomalies and store as list
anomalies <- list()
for (ssp in ssps) {
for (year in years) {
message(" Calculating anomaly: ", ssp, " ", year)
# Calculate ensemble mean future tmean
gcm_layers <- list()
for (g in gcms) {
tmin <- rast(paste0(out_dir_future, "/wc2.1_30s_tmin_", g, "_", "ssp", ssp, "_", year, "_tile-34.tif"))
tmax <- rast(paste0(out_dir_future, "/wc2.1_30s_tmax_", g, "_", "ssp", ssp, "_", year, "_tile-34.tif"))
tmin <- app(mask(crop(tmin, adm0), adm0), mean)
tmax <- app(mask(crop(tmax, adm0), adm0), mean)
gcm_layers[[g]] <- (tmin + tmax) / 2
}
fut_tmean <- app(rast(gcm_layers), mean)
# Store anomaly
anomalies[[paste0(ssp, "_", year)]] <- fut_tmean - tmean_annual
}
}
## Temperature change maps with shared colour scale across all SSP/period combinations
# Common colour scale across all panels (min to max)
anom_min <- Inf
anom_max <- 0
for (key in names(anomalies)) {
vals <- minmax(anomalies[[key]])
anom_min <- min(anom_min, vals[1])
anom_max <- max(anom_max, vals[2])
}
anom_range <- c(anom_min, anom_max)
print(anom_range) # e.g. c(1.4, 3.42)
## Plot 2x2 grid with shared legend and national mean annotation
fname <- paste0(out_dir_graphics, "/annual_temp_anomaly_2x2")
png(paste0(fname, ".png"), width = 12, height = 10, units = "in", res = 300, type = "cairo")
par(mfrow = c(2, 2))
for (ssp in ssps) {
for (year in years) {
key <- paste0(ssp, "_", year)
ssp_label <- ifelse(ssp == "245", "SSP2-4.5", "SSP5-8.5")
title <- paste0("Annual mean temperature change (°C)\n", ssp_label, " ", year, " relative to 1970\u20132000")
mean_val <- round(as.numeric(global(anomalies[[key]], mean, na.rm = TRUE)), 2)
plot(anomalies[[key]],
main = title,
mar = c(3.1, 3.1, 3.1, 5.1),
col = hcl.colors(100, "YlOrRd", rev = TRUE),
range = anom_range,
xlab = "Longitude (°E)",
ylab = "Latitude (°N)",
cex.main = 1.0,
cex.lab = 1.0,
legend = TRUE,
box = FALSE
)
lines(adm0, col = "black", lwd = 1.5)
# National mean annotation
mtext(paste0("National mean: +", mean_val, "\u00B0C"),
side = 3, line = 0, cex = 0.8, font = 2
)
}
}
dev.off()
## Check actual mean anomaly values for each panel
# for (key in names(anomalies)) {
# cat(key, ":", round(as.numeric(global(anomalies[[key]], mean, na.rm = TRUE)), 3), "°C\n")
# }
# 245_2041-2060 : 1.724 °C
# 245_2061-2080 : 2.232 °C
# 585_2041-2060 : 2.103 °C
# 585_2061-2080 : 3.239 °C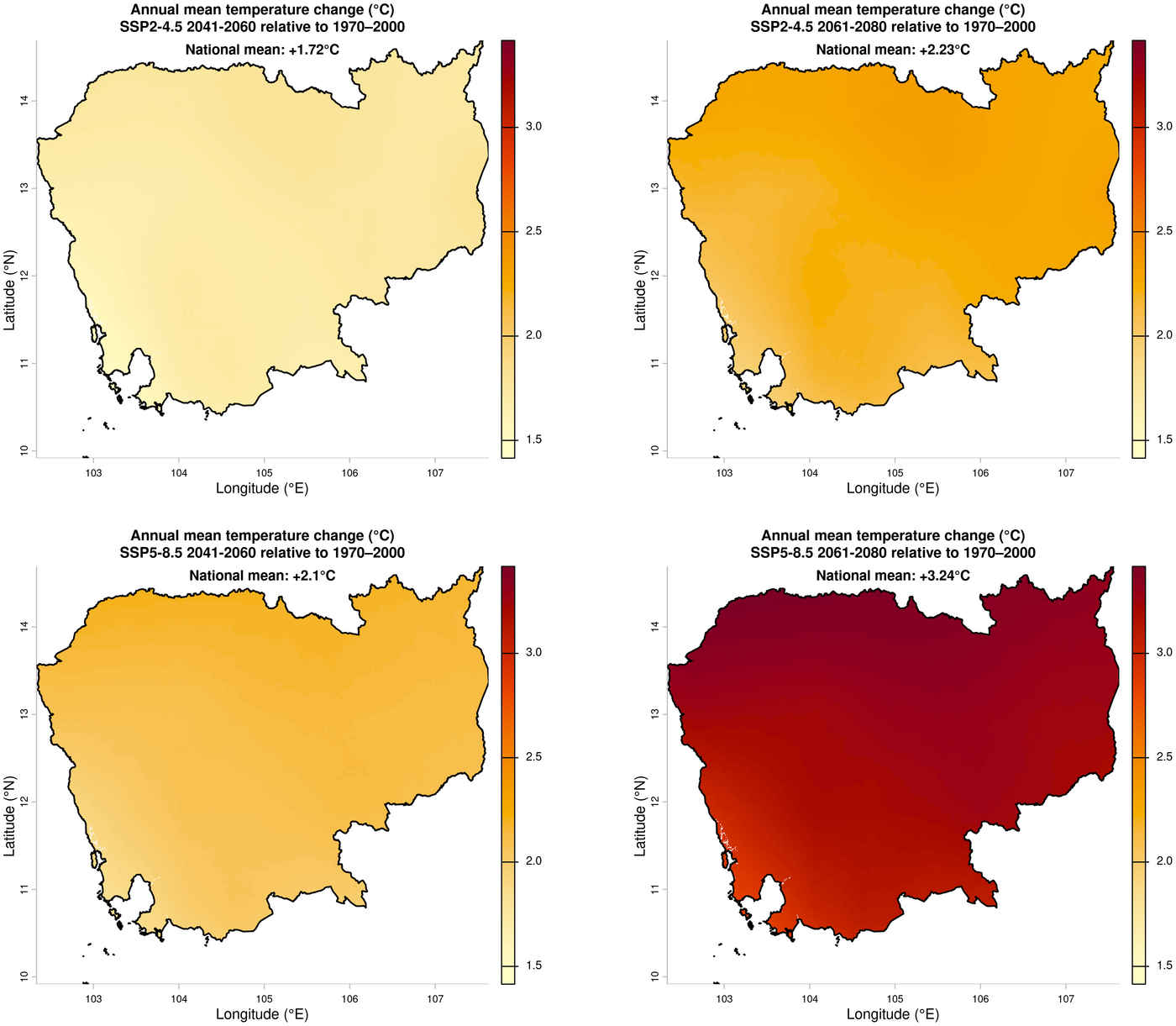
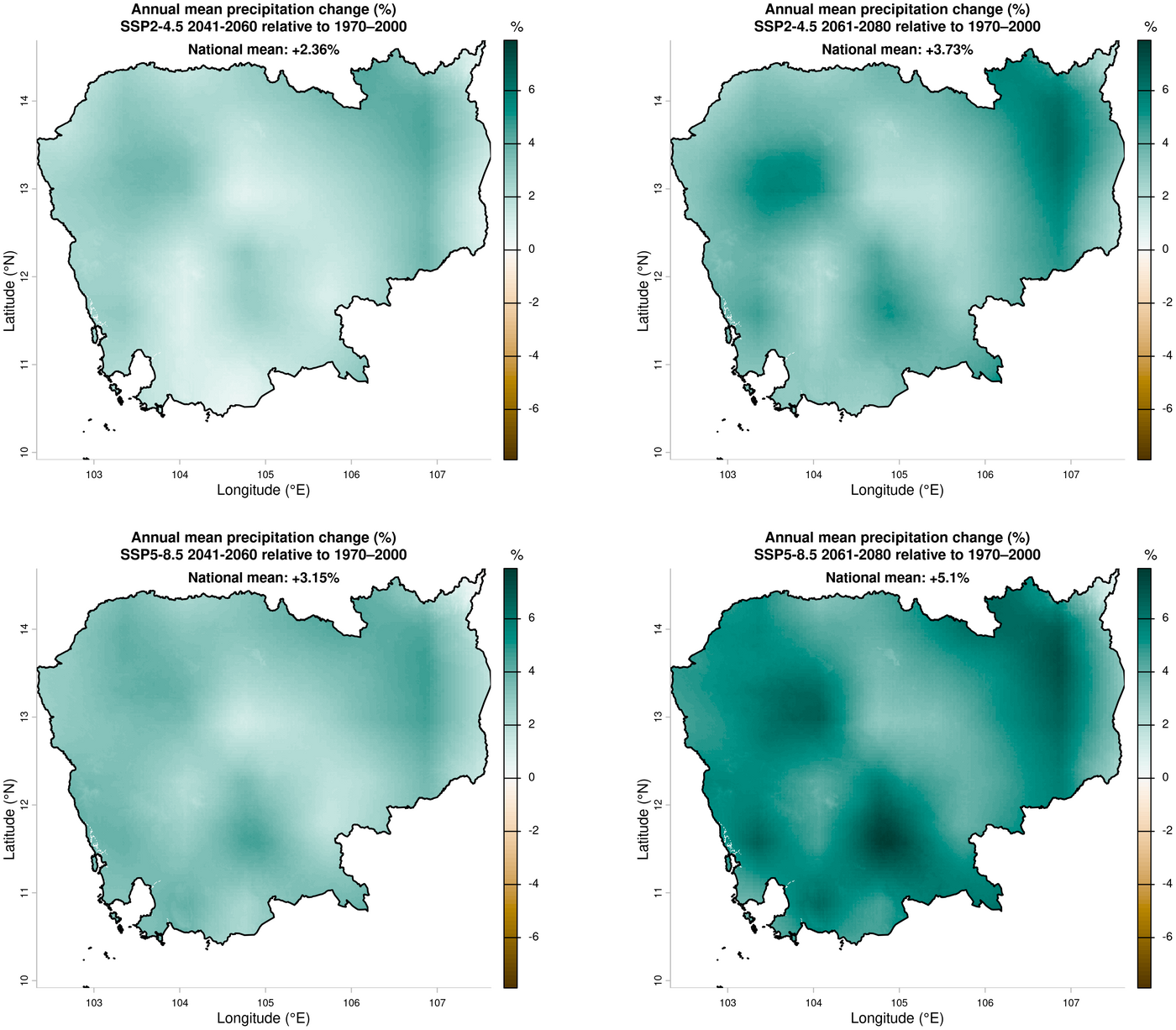
4. Annual mean precipitation change — ensemble mean relative to 1970–2000
Process:
- Read in precipitation values for each of the 5 GCMs (12 monthly layers each)
- Calculate mean total annual precipitation for each GCM using app(…, sum): output 5 separate layers
- Stack the 5 layers with rast(gcm_layers)
- Take the ensemble mean across the 5 GCMs with app(…, mean): output single layer
- Calculate percentage change from historical:((future - historical) / historical) * 100
# ============================================================
# Annual mean precipitation change — 2x2 grid (SSP x Year)
# Output: Spatial anomaly panel map
# ============================================================
# Calculate precipitation anomalies and store
prec_anomalies <- list()
for (ssp in ssps) {
for (year in years) {
message(" Calculating precipitation anomaly: ", ssp, " ", year)
# Calculate ensemble mean future total annual precipitation
gcm_layers <- list()
for (g in gcms) {
prec <- rast(paste0(out_dir_future, "/wc2.1_30s_prec_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
prec <- app(mask(crop(prec, adm0), adm0), sum)
gcm_layers[[g]] <- prec
}
fut_prec <- app(rast(gcm_layers), mean) # Calc ensemble mean
# Percentage change
prec_anomalies[[paste0(ssp, "_", year)]] <- ((fut_prec - prec_annual) / prec_annual) * 100
}
}
# Shared symmetric colour scale centred on zero
anom_abs_max <- 0
for (key in names(prec_anomalies)) {
anom_abs_max <- max(anom_abs_max, max(abs(minmax(prec_anomalies[[key]]))))
}
anom_range <- c(-anom_abs_max, anom_abs_max) # e.g. c(-7.9, 7.9)
# print(anom_range)
# Plot 2x2 grid
fname <- paste0(out_dir_graphics, "/annual_prec_anomaly_2x2")
png(paste0(fname, ".png"), width = 12, height = 10, units = "in", res = 300, type = "cairo")
par(mfrow = c(2, 2))
for (ssp in ssps) {
for (year in years) {
key <- paste0(ssp, "_", year)
ssp_label <- ifelse(ssp == "245", "SSP2-4.5", "SSP5-8.5")
title <- paste0("Annual mean precipitation change (%)\n", ssp_label, " ", year, " relative to 1970\u20132000")
mean_val <- round(as.numeric(global(prec_anomalies[[key]], mean, na.rm = TRUE)), 2)
plot(prec_anomalies[[key]],
main = title,
mar = c(3.1, 3.1, 3.1, 5.1),
col = hcl.colors(100, "BrBG"),
range = anom_range,
xlab = "Longitude (°E)",
ylab = "Latitude (°N)",
cex.main = 1.0,
cex.lab = 1.0,
plg = list(title = "%"),
box = FALSE
)
lines(adm0, col = "black", lwd = 1.5)
# National mean annotation
mtext(paste0("National mean: +", mean_val, "%"),
side = 3, line = 0, cex = 0.8, font = 2
)
}
}
dev.off()
# Check anomaly values:
# for (key in names(prec_anomalies)) {
# cat(key, ":", round(as.numeric(minmax(prec_anomalies[[key]])), 2), "\n")
# }
5. Annual mean temperature anomaly by GCM
# ============================================================
# Annual mean temperature anomaly by GCM
# Model ensemble spread plot: GCM spread per SSP and time period
# ============================================================
# Historical monthly tmean needed for anomaly baseline
hist_tmin <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_tmin.tif"))
hist_tmax <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_tmax.tif"))
adm0 <- geodata::gadm(country = "KHM", level = 0, path = out_dir)
hist_tmin_c <- mask(crop(hist_tmin, adm0), adm0)
hist_tmax_c <- mask(crop(hist_tmax, adm0), adm0)
tmin_monthly <- as.numeric(global(hist_tmin_c, mean, na.rm = TRUE)$mean)
tmax_monthly <- as.numeric(global(hist_tmax_c, mean, na.rm = TRUE)$mean)
tmean_monthly_hist <- (tmin_monthly + tmax_monthly) / 2
# Calculate annual mean temperature anomaly per GCM/SSP/year
gcm_anomalies <- list()
for (ssp in ssps) {
for (year in years) {
key <- paste0("ssp", ssp, "_", year)
gcm_anomalies[[key]] <- list(ssp = ssp, year = year, vals = numeric(length(gcms)))
names(gcm_anomalies[[key]]$vals) <- gcms
for (g in gcms) {
tmin <- rast(paste0(out_dir_future, "/wc2.1_30s_tmin_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
tmax <- rast(paste0(out_dir_future, "/wc2.1_30s_tmax_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
tmin <- mask(crop(tmin, adm0), adm0)
tmax <- mask(crop(tmax, adm0), adm0)
fut_tmean <- mean(as.numeric(global(tmin, mean, na.rm = TRUE)$mean) +
as.numeric(global(tmax, mean, na.rm = TRUE)$mean)) / 2
gcm_anomalies[[key]]$vals[g] <- fut_tmean - mean(tmean_monthly_hist)
}
}
}
# Set up plot
all_vals <- unlist(lapply(gcm_anomalies, function(x) x$vals))
xlim <- c(0, ceiling(max(all_vals) * 2) / 2)
n_groups <- length(gcm_anomalies)
ylim <- c(0.5, n_groups + 1)
group_labels <- c(
"SSP2-4.5 2041\u20132060",
"SSP2-4.5 2061\u20132080",
"SSP5-8.5 2041\u20132060",
"SSP5-8.5 2061\u20132080"
)
ssp_cols <- list("245" = "#EAB139", "585" = "#980002")
gcm_pch <- c(16, 17, 15, 18, 8)
names(gcm_pch) <- gcms
# Plot
fname <- paste0(out_dir_graphics, "/annual_mean_temp_anomaly_gcm")
png(paste0(fname, ".png"), width = 10, height = 6, units = "in", res = 300, type = "cairo")
par(mar = c(4, 9, 4, 2), mgp = c(2.5, 0.5, 0))
plot(NA,
xlim = xlim,
ylim = ylim,
xlab = "Temperature anomaly (\u00B0C) relative to 1970\u20132000",
ylab = "",
yaxt = "n",
main = "Annual mean temperature anomaly by GCM\nCambodia \u2014 future projections",
bty = "n",
cex.main = 0.9
)
# Grid lines and zero reference
abline(
v = seq(xlim[1], xlim[2], by = 0.5),
col = "grey90", lty = 1, lwd = 0.8
)
abline(v = 0, col = "grey30", lty = 2, lwd = 1.5)
# Plot dots, ranges, and means per group
for (i in seq_along(gcm_anomalies)) {
key <- names(gcm_anomalies)[i]
vals <- gcm_anomalies[[key]]$vals
ssp <- gcm_anomalies[[key]]$ssp
col <- ssp_cols[[ssp]]
ens_mean <- mean(vals)
# Range bar
segments(min(vals), i, max(vals), i,
col = adjustcolor(col, alpha.f = 0.4), lwd = 3
)
# Ensemble mean — vertical line
segments(ens_mean, i - 0.2,
ens_mean, i + 0.2,
col = col, lwd = 3
)
# Ensemble mean value above the vertical line
text(ens_mean, i + 0.35,
labels = paste0("+", round(ens_mean, 2), "\u00B0C"),
col = col,
cex = 0.75,
font = 2
)
# Individual GCM dots
for (g in gcms) {
points(vals[g], i,
pch = gcm_pch[g],
col = col,
cex = 1.2
)
}
}
# Y axis labels
axis(2, at = 1:n_groups, labels = group_labels, las = 1, cex.axis = 0.85, tcl = 0)
# Legend for GCMs
legend("bottomright",
legend = c(gcms, "Ensemble mean (\u275A)"),
pch = c(gcm_pch, NA),
col = "grey40",
bty = "n",
cex = 0.75,
title = "GCM",
title.font = 2
)
dev.off()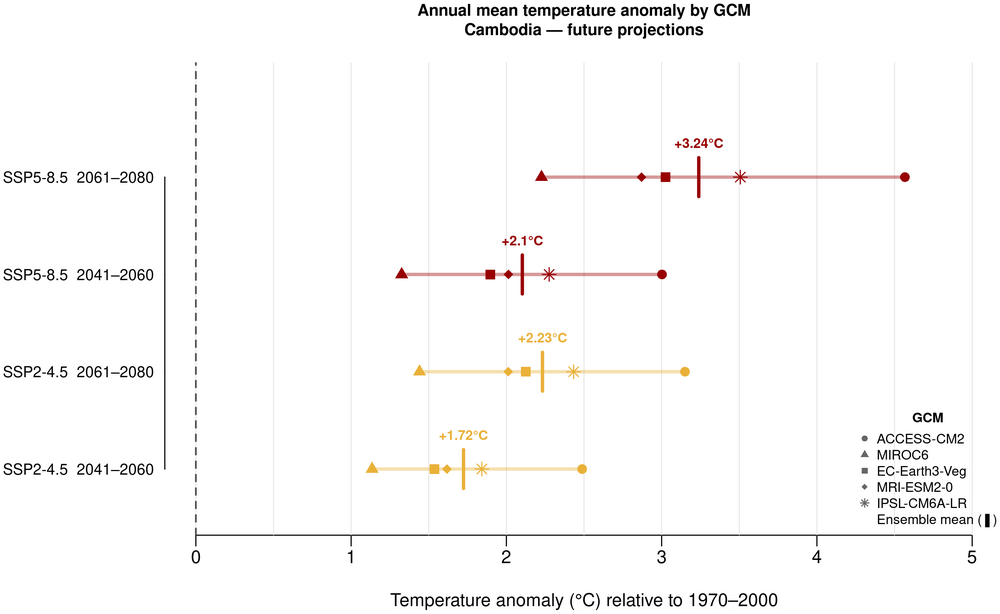
6. Annual mean precipitation anomaly by GCM
# ============================================================
# Annual mean precipitation anomaly by GCM
# Model ensemble spread plot: GCM spread per SSP and time period
# ============================================================
# Historical monthly precipitation needed for anomaly baseline
hist_prec <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_prec.tif"))
adm0 <- geodata::gadm(country = "KHM", level = 0, path = out_dir)
hist_prec_c <- mask(crop(hist_prec, adm0), adm0)
prec_monthly <- as.numeric(global(hist_prec_c, mean, na.rm = TRUE)$mean)
# Calculate annual mean precipitation anomaly per GCM/SSP/year
gcm_anomalies <- list()
for (ssp in ssps) {
for (year in years) {
key <- paste0("ssp", ssp, "_", year)
gcm_anomalies[[key]] <- list(ssp = ssp, year = year, vals = numeric(length(gcms)))
names(gcm_anomalies[[key]]$vals) <- gcms
for (g in gcms) {
prec <- rast(paste0(out_dir_future, "/wc2.1_30s_prec_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
prec <- mask(crop(prec, adm0), adm0)
# Annual total precipitation across Cambodia
fut_prec <- sum(as.numeric(global(prec, mean, na.rm = TRUE)$mean))
hist_prec <- sum(prec_monthly)
gcm_anomalies[[key]]$vals[g] <- ((fut_prec - hist_prec) / hist_prec) * 100
}
}
}
# Set up plot
all_vals <- unlist(lapply(gcm_anomalies, function(x) x$vals))
xlim <- c(floor(min(all_vals)) - 1, ceiling(max(all_vals)) + 1)
n_groups <- length(gcm_anomalies)
ylim <- c(0.5, n_groups + 1)
group_labels <- c(
"SSP2-4.5 2041\u20132060",
"SSP2-4.5 2061\u20132080",
"SSP5-8.5 2041\u20132060",
"SSP5-8.5 2061\u20132080"
)
ssp_cols <- list("245" = "#EAB139", "585" = "#980002")
gcm_pch <- c(16, 17, 15, 18, 8)
names(gcm_pch) <- gcms
## Plot
fname <- paste0(out_dir_graphics, "/annual_mean_prec_anomaly_gcm")
png(paste0(fname, ".png"), width = 10, height = 6, units = "in", res = 300, type = "cairo")
par(mar = c(4, 9, 4, 2), mgp = c(2.5, 0.5, 0))
plot(NA,
xlim = xlim,
ylim = ylim,
xlab = "Precipitation anomaly (%) relative to 1970\u20132000",
ylab = "",
yaxt = "n",
main = "Annual mean precipitation anomaly by GCM\nCambodia \u2014 future projections",
bty = "n",
cex.main = 0.9
)
# Grid lines and zero reference
abline(
v = seq(floor(xlim[1]), ceiling(xlim[2]), by = 2),
col = "grey90", lty = 1, lwd = 0.8
)
abline(v = 0, col = "grey30", lty = 2, lwd = 1.5)
# Plot dots, ranges, and means per group
for (i in seq_along(gcm_anomalies)) {
key <- names(gcm_anomalies)[i]
vals <- gcm_anomalies[[key]]$vals
ssp <- gcm_anomalies[[key]]$ssp
col <- ssp_cols[[ssp]]
ens_mean <- mean(vals)
# Range bar
segments(min(vals), i, max(vals), i,
col = adjustcolor(col, alpha.f = 0.4), lwd = 3
)
# Ensemble mean — vertical line
segments(ens_mean, i - 0.2,
ens_mean, i + 0.2,
col = col, lwd = 3
)
# Ensemble mean value above the vertical line
text(ens_mean, i + 0.35,
labels = paste0(ifelse(ens_mean >= 0, "+", ""), round(ens_mean, 1), "%"),
col = col,
cex = 0.75,
font = 2
)
# Individual GCM dots
for (g in gcms) {
points(vals[g], i,
pch = gcm_pch[g],
col = col,
cex = 1.2
)
}
}
# Y axis labels
axis(2, at = 1:n_groups, labels = group_labels, las = 1, cex.axis = 0.85, tcl = 0)
# Legend for GCMs
legend("bottomright",
legend = c(gcms, "Ensemble mean (\u275A)"),
pch = c(gcm_pch, NA),
col = "grey40",
bty = "n",
cex = 0.75,
title = "GCM",
title.font = 2
)
dev.off()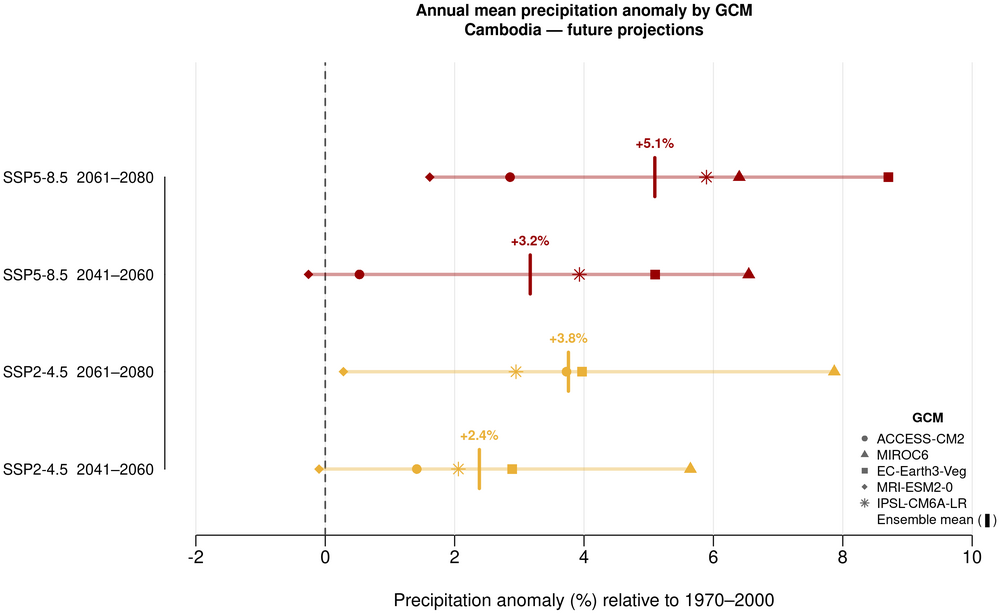
Monthly maps and graphics
The following can be applied to generate monthly maps and graphics for any region of the world.
1. Monthly mean temperature and precipitation — historical baseline (1970–2000)
# ============================================================
# Monthly mean precipitation and temperature — historical (1970–2000)
# ============================================================
# Read in historical data
hist_prec <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_prec.tif"))
hist_tmin <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_tmin.tif"))
hist_tmax <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_tmax.tif"))
# Crop to Cambodia boundary
adm0 <- geodata::gadm(country = "KHM", level = 0, path = out_dir)
hist_prec_c <- mask(crop(hist_prec, adm0), adm0)
hist_tmin_c <- mask(crop(hist_tmin, adm0), adm0)
hist_tmax_c <- mask(crop(hist_tmax, adm0), adm0)
# Monthly spatial means
prec_monthly <- as.numeric(global(hist_prec_c, mean, na.rm = TRUE)$mean)
tmin_monthly <- as.numeric(global(hist_tmin_c, mean, na.rm = TRUE)$mean)
tmax_monthly <- as.numeric(global(hist_tmax_c, mean, na.rm = TRUE)$mean)
tmean_monthly_hist <- (tmin_monthly + tmax_monthly) / 2
# Scaling: map temperature onto precipitation axis
# Temperature range: ~24-30°C; Precipitation range: 0-320mm
# Scale temp to fit within precipitation axis range
prec_max <- max(prec_monthly) * 1.15
temp_min <- floor(min(tmean_monthly_hist)) - 1 # e.g. 23
temp_max <- ceiling(max(tmean_monthly_hist)) + 1 # e.g. 31
temp_range <- temp_max - temp_min
# Linear scaling function: temp -> prec axis
temp_to_prec <- function(t) (t - temp_min) / temp_range * prec_max
month_labels <- c(
"Jan", "Feb", "Mar", "Apr", "May", "Jun",
"Jul", "Aug", "Sep", "Oct", "Nov", "Dec"
)
## Plot
fname <- paste0(out_dir_graphics, "/monthly_mean_prec_temp_historical")
png(paste0(fname, ".png"), width = 8, height = 6, units = "in", res = 300, type = "cairo")
par(mar = c(5, 4, 4, 4), mgp = c(3, 0.3, 0))
# Precipitation bars
bp <- barplot(prec_monthly,
names.arg = month_labels,
col = adjustcolor("steelblue", alpha.f = 0.7),
border = NA,
ylim = c(0, prec_max),
ylab = "",
xlab = "",
main = "Monthly mean precipitation and temperature\nCambodia \u2014 historical (1970\u20132000)",
las = 1,
cex.names = 0.9,
cex.axis = 0.8,
tcl = -0.2
)
# Left y-axis title
mtext("Mean precipitation (mm)", side = 2, line = 1.8, cex = 0.8)
# Temperature line (scaled to precipitation axis)
lines(bp, temp_to_prec(tmean_monthly_hist), col = "firebrick", lwd = 2.5)
points(bp, temp_to_prec(tmean_monthly_hist), col = "firebrick", pch = 16, cex = 0.8)
# Right y-axis: temperature
temp_ticks <- seq(temp_min, temp_max, by = 2)
axis(4,
at = temp_to_prec(temp_ticks),
labels = temp_ticks,
las = 1,
cex.axis = 0.8,
tcl = -0.2
)
mtext("Mean temperature (\u00B0C)", side = 4, line = 1.8, cex = 0.8)
# Precipitation value labels
text(bp, prec_monthly + 8,
labels = round(prec_monthly, 0),
cex = 0.75,
col = "grey30"
)
# Annual total annotation
annual_total <- sum(prec_monthly)
mtext(paste0("Mean annual total: ", round(annual_total, 0), " mm"),
side = 1, line = 2.5, cex = 0.85, font = 2
)
# Legend
legend("topleft",
legend = c("Mean precipitation", "Mean temperature"),
fill = c(adjustcolor("steelblue", alpha.f = 0.7), NA),
border = c(NA, NA),
lwd = c(NA, 2.5),
col = c(NA, "firebrick"),
pch = c(NA, 16),
bty = "n",
cex = 0.8
)
dev.off()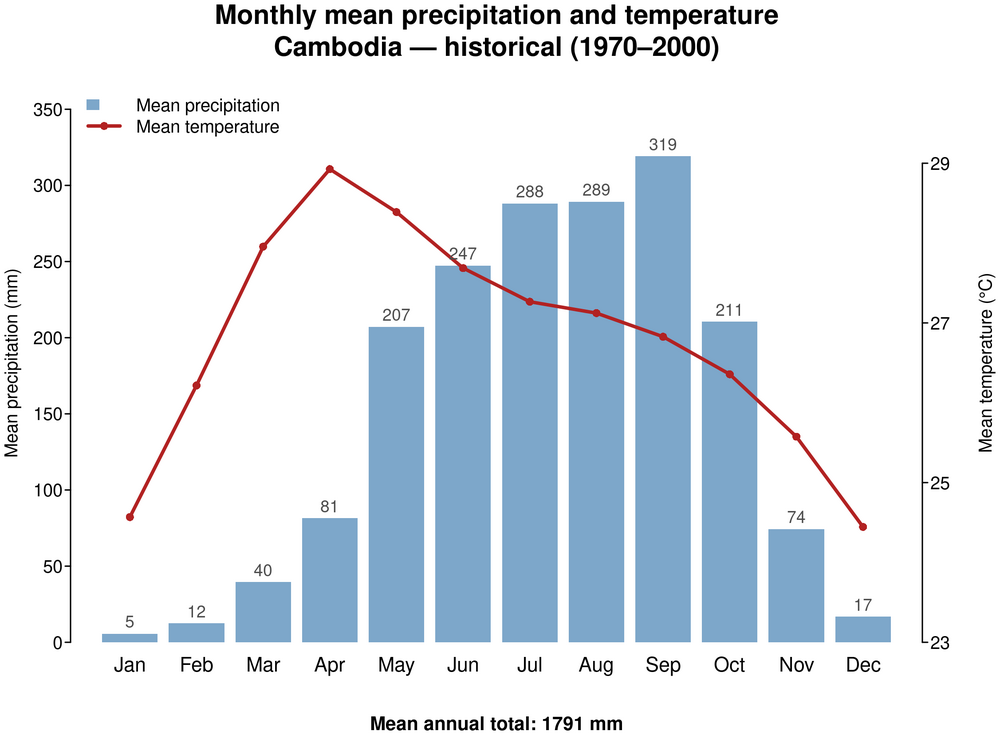
2. Monthly mean temperature — historical and future projections
Historical line + SSP2-4.5 and SSP5-8.5 ribbons (GCM spread) in two panels for 2041-2060 and 2061-2080
# ============================================================
# Monthly mean temperature: Ensemble range plot
# Historical + SSP2-4.5 and SSP5-8.5 ribbons (GCM spread)
# Two panels: 2041-2060 and 2061-2080
# ============================================================
# Read in historical data
hist_tmin <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_tmin.tif"))
hist_tmax <- rast(paste0(out_dir_historical, "/tile_34_wc2.1_30s_tmax.tif"))
# Crop to Cambodia boundary
adm0 <- geodata::gadm(country = "KHM", level = 0, path = out_dir)
hist_tmin_c <- mask(crop(hist_tmin, adm0), adm0)
hist_tmax_c <- mask(crop(hist_tmax, adm0), adm0)
# Annual spatial rasters (for mapping)
tmin_annual <- app(hist_tmin_c, mean)
tmax_annual <- app(hist_tmax_c, mean)
tmean_annual <- (tmin_annual + tmax_annual) / 2
# Monthly spatial means across Cambodia (for seasonal cycle plots)
tmin_monthly <- as.numeric(global(hist_tmin_c, mean, na.rm = TRUE)$mean)
tmax_monthly <- as.numeric(global(hist_tmax_c, mean, na.rm = TRUE)$mean)
tmean_monthly_hist <- (tmin_monthly + tmax_monthly) / 2
# Set up plot
month_labels <- c(
"Jan", "Feb", "Mar", "Apr", "May", "Jun",
"Jul", "Aug", "Sep", "Oct", "Nov", "Dec"
)
mon <- 1:12
fname <- paste0(out_dir_graphics, "/monthly_mean_temperature_hist_future")
## Plot
png(paste0(fname, ".png"), width = 12, height = 6, units = "in", res = 300, type = "cairo")
par(mfrow = c(1, 2), mar = c(4, 3.5, 4, 2), mgp = c(1.8, 0.5, 0))
for (year in years) {
# Calculate ensemble stats for each SSP
ens <- list()
for (ssp in ssps) {
gcm_monthly <- list()
for (g in gcms) {
tmin <- rast(paste0(out_dir_future, "/wc2.1_30s_tmin_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
tmax <- rast(paste0(out_dir_future, "/wc2.1_30s_tmax_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
tmin <- mask(crop(tmin, adm0), adm0)
tmax <- mask(crop(tmax, adm0), adm0)
tmin_m <- as.numeric(global(tmin, mean, na.rm = TRUE)$mean)
tmax_m <- as.numeric(global(tmax, mean, na.rm = TRUE)$mean)
gcm_monthly[[g]] <- (tmin_m + tmax_m) / 2
}
mat <- do.call(rbind, gcm_monthly) # 5 x 12 matrix
ens[[ssp]] <- list(
mean = colMeans(mat),
min = apply(mat, 2, min),
max = apply(mat, 2, max)
)
}
# Y-axis range
ylim <- range(c(
tmean_monthly_hist,
ens[["245"]]$min, ens[["245"]]$max,
ens[["585"]]$min, ens[["585"]]$max
)) + c(-0.5, 0.5)
# Set up empty plot
plot(mon, tmean_monthly_hist,
type = "n",
xlab = "",
ylab = "Mean temperature (\u00B0C)",
xaxt = "n",
yaxt = "n",
ylim = ylim,
main = paste0("Monthly mean temperature\nCambodia \u2014 ", "historical and future projections ", "(", year, ")"),
bty = "n",
cex.main = 1.0
)
axis(1, at = mon, labels = month_labels, cex.axis = 0.85, tcl = -0.2)
axis(2, las = 1, cex.axis = 0.8)
# Ribbons and lines per SSP
ssp_cols <- list(
"245" = "#EAB139", # SSP2-4.5 gold/yellow
"585" = "#980002" # SSP5-8.5 dark red
)
for (ssp in ssps) {
polygon(c(mon, rev(mon)),
c(ens[[ssp]]$max, rev(ens[[ssp]]$min)),
col = adjustcolor(ssp_cols[[ssp]], alpha.f = 0.2),
border = NA
)
}
for (ssp in ssps) {
lines(mon, ens[[ssp]]$mean,
col = ssp_cols[[ssp]], lwd = 2, lty = 2
)
}
lines(mon, tmean_monthly_hist, col = "black", lwd = 2.5)
# Legend
legend("bottomleft",
inset = c(0.1, 0.02),
xpd = FALSE, # move right by 10%, no vertical inset
legend = c(
"Historical (1970\u20132000)",
"SSP2-4.5 ensemble mean", "SSP2-4.5 GCM range",
"SSP5-8.5 ensemble mean", "SSP5-8.5 GCM range"
),
col = c(
"black",
"#EAB139", adjustcolor("#EAB139", alpha.f = 0.2),
"#980002", adjustcolor("#980002", alpha.f = 0.2)
),
lwd = c(2.5, 2, 8, 2, 8),
lty = c(1, 2, 1, 2, 1),
bty = "n",
cex = 0.75
)
}
dev.off()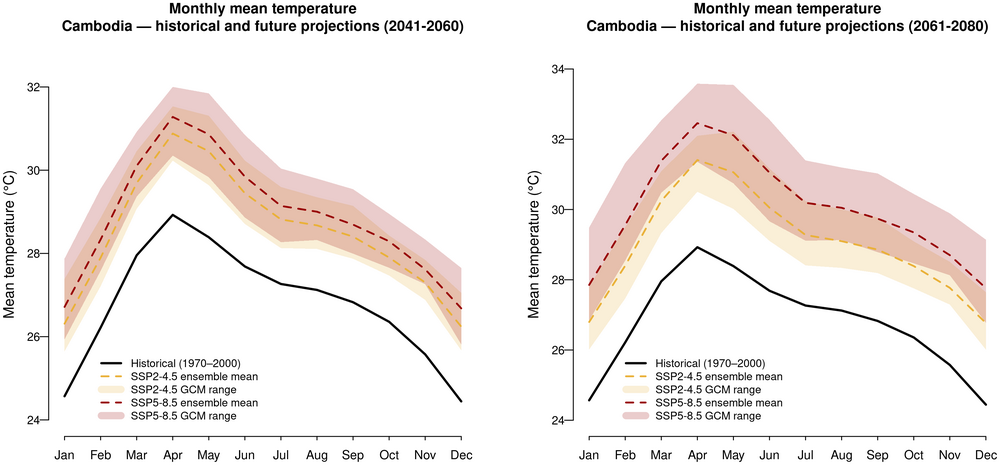
3. Monthly mean precipitation — historical and future projections
Historical line + SSP2-4.5 and SSP5-8.5 ribbons (GCM spread) in two panels for 2041-2060 and 2061-2080
# ============================================================
# Monthly mean precipitation: Ensemble range plot
# Historical + SSP2-4.5 and SSP5-8.5 ribbons (GCM spread)
# Two panels: 2041-2060 and 2061-2080
# ============================================================
## Plot
fname <- paste0(out_dir_graphics, "/monthly_mean_prec_hist_future")
png(paste0(fname, ".png"), width = 12, height = 6, units = "in", res = 300, type = "cairo")
par(mfrow = c(1, 2), mar = c(4, 3.5, 4, 2), mgp = c(1.8, 0.5, 0))
# Calculate ensemble stats for each SSP
for (year in years) {
ens <- list()
for (ssp in ssps) {
gcm_monthly <- list()
for (g in gcms) {
prec <- rast(paste0(out_dir_future, "/wc2.1_30s_prec_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
prec <- mask(crop(prec, adm0), adm0)
prec_m <- as.numeric(global(prec, mean, na.rm = TRUE)$mean) # 12 monthly values
gcm_monthly[[g]] <- prec_m
}
mat <- do.call(rbind, gcm_monthly) # 5 x 12 matrix
ens[[ssp]] <- list(
mean = colMeans(mat),
min = apply(mat, 2, min),
max = apply(mat, 2, max)
)
}
# Y-axis range
ylim <- range(c(
prec_monthly,
ens[["245"]]$min, ens[["245"]]$max,
ens[["585"]]$min, ens[["585"]]$max
)) + c(-10, 20)
# Set up empty plot
plot(mon, prec_monthly,
type = "n",
xlab = "",
ylab = "Precipitation (mm)",
xaxt = "n",
yaxt = "n",
ylim = ylim,
main = paste0("Monthly mean precipitation\nCambodia \u2014 ", "historical and future projections ", "(", year, ")"),
bty = "n",
cex.main = 1.0
)
axis(1, at = mon, labels = month_labels, cex.axis = 0.85, tcl = -0.2)
axis(2, las = 1, cex.axis = 0.8)
# Ribbons and lines per SSP
ssp_cols <- list(
"245" = "#EAB139", # SSP2-4.5 gold/yellow
"585" = "#980002" # SSP5-8.5 dark red
)
for (ssp in ssps) {
polygon(c(mon, rev(mon)),
c(ens[[ssp]]$max, rev(ens[[ssp]]$min)),
col = adjustcolor(ssp_cols[[ssp]], alpha.f = 0.2),
border = NA
)
}
for (ssp in ssps) {
lines(mon, ens[[ssp]]$mean,
col = ssp_cols[[ssp]], lwd = 2, lty = 2
)
}
lines(mon, prec_monthly, col = "black", lwd = 2.5)
# Legend
legend("topleft",
inset = c(0.1, 0.02),
xpd = FALSE,
legend = c(
"Historical (1970\u20132000)",
"SSP2-4.5 ensemble mean", "SSP2-4.5 GCM range",
"SSP5-8.5 ensemble mean", "SSP5-8.5 GCM range"
),
col = c(
"black",
"#EAB139", adjustcolor("#EAB139", alpha.f = 0.2),
"#980002", adjustcolor("#980002", alpha.f = 0.2)
),
lwd = c(2.5, 2, 8, 2, 8),
lty = c(1, 2, 1, 2, 1),
bty = "n",
cex = 0.75
)
}
dev.off()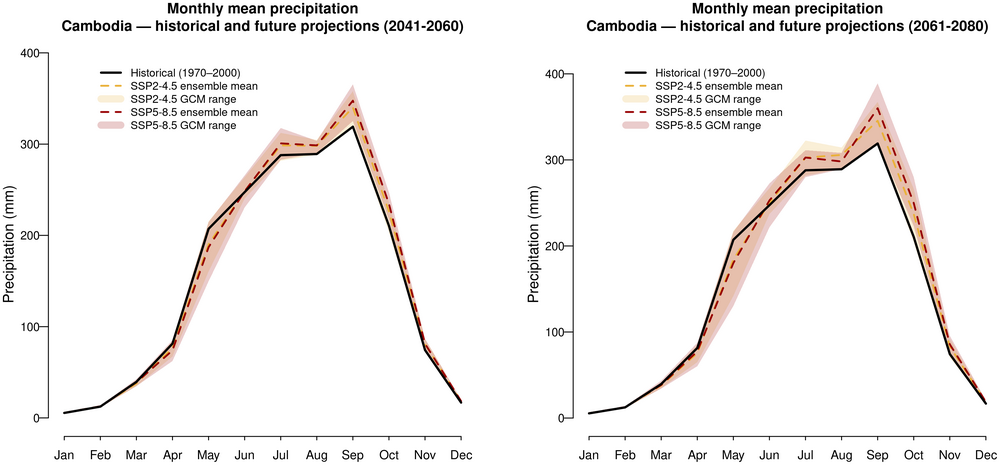
Seasonal maps and graphics
These outputs are tailored to Cambodia and rice cultivation.
1. Wet season precipitation change — ensemble mean relative to 1970–2000
# ============================================================
# Change in wet season precipitation (%)
# Future relative to historical (1970-2000)
# Wet season: May-October (layers 5-10)
# 2x2 grid: SSP2-4.5/SSP5-8.5 x 2041-2060/2061-2080
# ============================================================
# Historical wet season precipitation
# Layers 5-10 = May-October
hist_prec_ws <- subset(hist_prec_c, 5:10)
# Total wet season prec per pixel
hist_prec_ws_r <- app(hist_prec_ws, sum)
# Shared colour scale
ws_anom_abs_max <- 0
ws_anomalies <- list()
for (ssp in ssps) {
for (year in years) {
key <- paste0(ssp, "_", year)
# Load and process future GCM files
gcm_layers <- list()
for (g in gcms) {
prec <- rast(paste0(out_dir_future, "/wc2.1_30s_prec_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
prec <- mask(crop(prec, adm0), adm0)
# Wet season layers only (May-Oct = layers 5-10)
prec_ws <- app(subset(prec, 5:10), sum)
gcm_layers[[g]] <- prec_ws
}
# Ensemble mean across GCMs
fut_ws <- app(rast(gcm_layers), mean)
# Percentage change relative to historical
ws_anomalies[[key]] <- ((fut_ws - hist_prec_ws_r) / hist_prec_ws_r) * 100
# Track max absolute value for symmetric colour scale
ws_anom_abs_max <- max(ws_anom_abs_max, max(abs(minmax(ws_anomalies[[key]]))))
}
}
anom_range <- c(-ws_anom_abs_max, ws_anom_abs_max)
# Plot 2x2 grid
fname <- paste0(out_dir_graphics, "/wet_season_prec_change_2x2")
png(paste0(fname, ".png"), width = 12, height = 10, units = "in", res = 300, type = "cairo")
par(mfrow = c(2, 2))
ssp_labels <- list("245" = "SSP2-4.5", "585" = "SSP5-8.5")
for (ssp in ssps) {
for (year in years) {
key <- paste0(ssp, "_", year)
anom <- ws_anomalies[[key]]
mean_val <- round(global(anom, mean, na.rm = TRUE)$mean, 2)
sign_val <- ifelse(mean_val >= 0, "+", "")
plot(anom,
main = paste0(
"Wet season precipitation change (%)\n",
ssp_labels[[ssp]], " ", year, " relative to 1970\u20132000"
),
col = hcl.colors(100, "BrBG"),
range = anom_range,
mar = c(3.5, 3.5, 3.5, 3.5),
plg = list(title = "%"),
xlab = "Longitude (\u00B0E)",
ylab = "Latitude (\u00B0N)",
cex.lab = 1.0,
cex.main = 1.0,
box = FALSE
)
lines(adm0, col = "black", lwd = 1.2)
mtext(paste0("National mean: ", sign_val, mean_val, "%"),
side = 3, line = -0.5, cex = 0.8, font = 2
)
}
}
dev.off()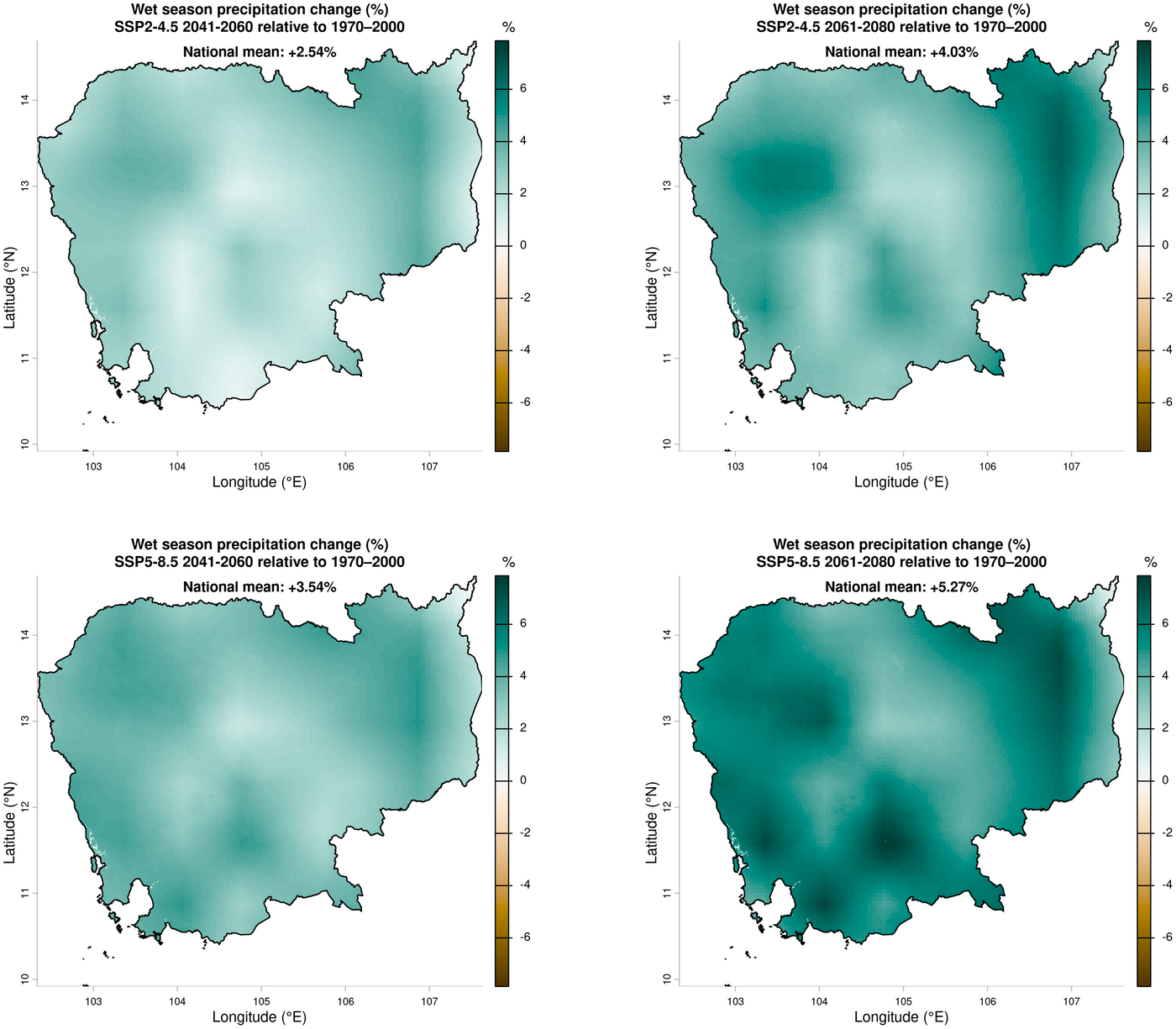
2. Dry season precipitation change — ensemble mean relative to 1970–2000
# ============================================================
# Change in dry season precipitation (%)
# Future relative to historical (1970-2000)
# Dry season: November-April (layers 11-12 and 1-4)
# 2x2 grid: SSP2-4.5/SSP5-8.5 x 2041-2060/2061-2080
# ============================================================
# Historical dry season precipitation
# Dry season spans two calendar years: Nov-Dec (layers 11-12) + Jan-Apr (layers 1-4)
hist_prec_ds <- subset(hist_prec_c, c(11, 12, 1, 2, 3, 4))
hist_prec_ds_r <- app(hist_prec_ds, sum) # total dry season precip per pixel
# Shared colour scale
ds_anom_abs_max <- 0
ds_anomalies <- list()
for (ssp in ssps) {
for (year in years) {
key <- paste0(ssp, "_", year)
gcm_layers <- list()
for (g in gcms) {
prec <- rast(paste0(out_dir_future, "/wc2.1_30s_prec_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
prec <- mask(crop(prec, adm0), adm0)
# Dry season layers: Nov-Dec (11-12) + Jan-Apr (1-4)
prec_ds <- app(subset(prec, c(11, 12, 1, 2, 3, 4)), sum)
gcm_layers[[g]] <- prec_ds
}
# Ensemble mean across GCMs
fut_ds <- app(rast(gcm_layers), mean)
# Percentage change relative to historical
ds_anomalies[[key]] <- ((fut_ds - hist_prec_ds_r) / hist_prec_ds_r) * 100
# Track max absolute value for symmetric colour scale
ds_anom_abs_max <- max(ds_anom_abs_max, max(abs(minmax(ds_anomalies[[key]]))))
}
}
anom_range <- c(-ds_anom_abs_max, ds_anom_abs_max)
# Plot 2x2 grid
fname <- paste0(out_dir_graphics, "/dry_season_prec_change_2x2")
png(paste0(fname, ".png"), width = 12, height = 10, units = "in", res = 300, type = "cairo")
par(mfrow = c(2, 2))
ssp_labels <- list("245" = "SSP2-4.5", "585" = "SSP5-8.5")
for (ssp in ssps) {
for (year in years) {
key <- paste0(ssp, "_", year)
anom <- ds_anomalies[[key]]
mean_val <- round(global(anom, mean, na.rm = TRUE)$mean, 2)
sign_val <- ifelse(mean_val >= 0, "+", "")
plot(anom,
main = paste0(
"Dry season precipitation change (%)\n",
ssp_labels[[ssp]], " ", year, " relative to 1970\u20132000"
),
col = hcl.colors(100, "BrBG"),
range = anom_range,
mar = c(3.5, 3.5, 3.5, 3.5),
xlab = "Longitude (\u00B0E)",
ylab = "Latitude (\u00B0N)",
plg = list(title = "%"),
cex.main = 1.0,
cex.lab = 1.0,
box = FALSE
)
lines(adm0, col = "black", lwd = 1.2)
mtext(paste0("National mean: ", sign_val, mean_val, "%"),
side = 3, line = -0.5, cex = 0.8, font = 2
)
}
}
dev.off()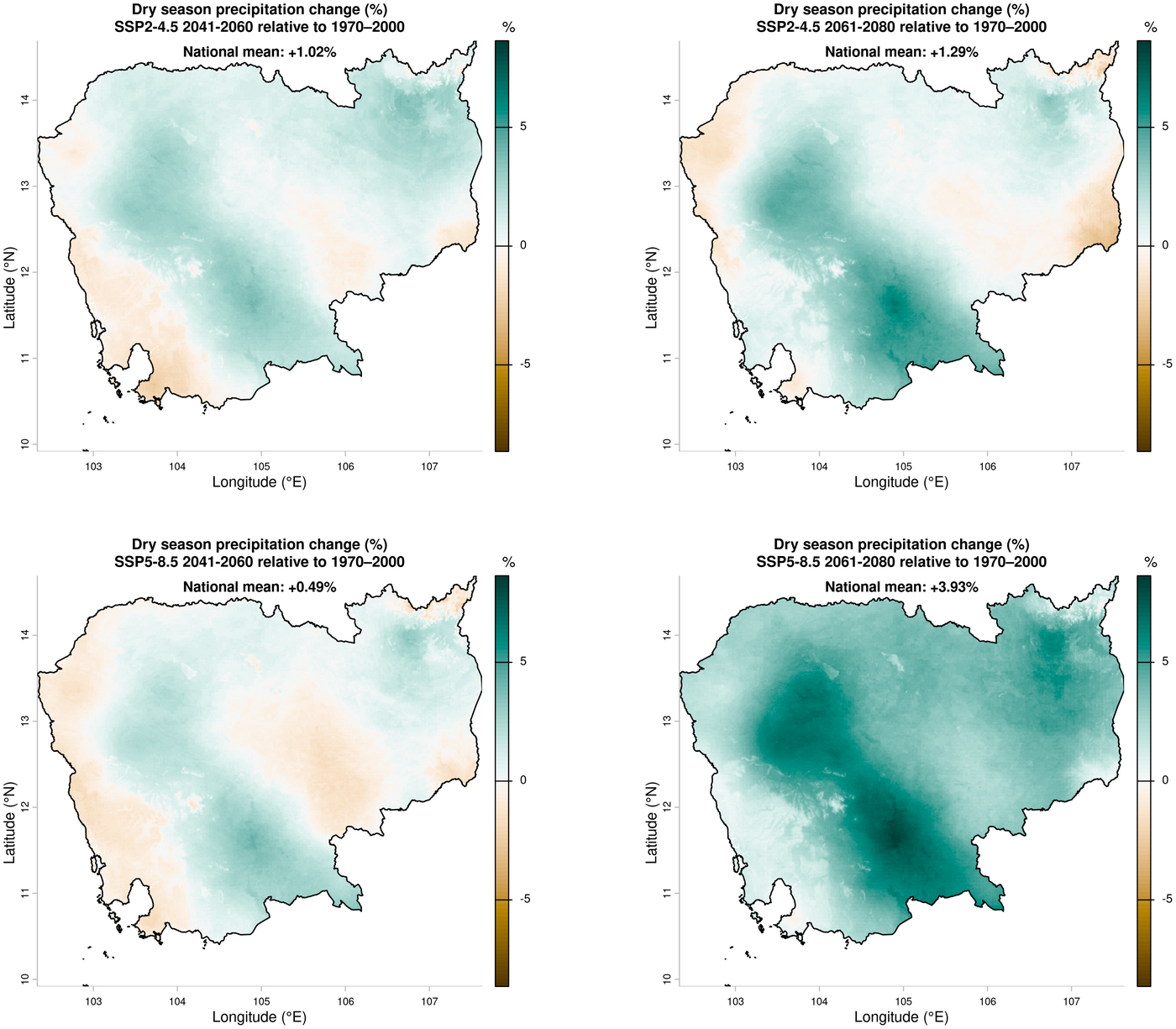
3. Change in dry season mean temperature (ensemble mean future minus historical)
# ============================================================
# Change in dry season mean temperature (°C)
# Future minus historical (1970-2000)
# Dry season: November-April (layers 11-12 and 1-4)
# 2x2 grid: SSP2-4.5/SSP5-8.5 x 2041-2060/2061-2080
# ============================================================
# Historical dry season mean temperature
hist_tmin_ds <- subset(hist_tmin_c, c(11, 12, 1, 2, 3, 4))
hist_tmax_ds <- subset(hist_tmax_c, c(11, 12, 1, 2, 3, 4))
hist_tmean_ds_r <- (app(hist_tmin_ds, mean) + app(hist_tmax_ds, mean)) / 2
# Shared colour scale
ds_temp_abs_max <- 0
ds_temp_anomalies <- list()
for (ssp in ssps) {
for (year in years) {
key <- paste0(ssp, "_", year)
gcm_layers <- list()
for (g in gcms) {
tmin <- rast(paste0(out_dir_future, "/wc2.1_30s_tmin_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
tmax <- rast(paste0(out_dir_future, "/wc2.1_30s_tmax_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
tmin <- mask(crop(tmin, adm0), adm0)
tmax <- mask(crop(tmax, adm0), adm0)
# Dry season layers: Nov-Dec (11-12) + Jan-Apr (1-4)
tmin_ds <- app(subset(tmin, c(11, 12, 1, 2, 3, 4)), mean)
tmax_ds <- app(subset(tmax, c(11, 12, 1, 2, 3, 4)), mean)
gcm_layers[[g]] <- (tmin_ds + tmax_ds) / 2
}
# Ensemble mean across GCMs
fut_ds_tmean <- app(rast(gcm_layers), mean)
# Temperature anomaly (absolute change in °C)
ds_temp_anomalies[[key]] <- fut_ds_tmean - hist_tmean_ds_r
# Track max absolute value for colour scale
ds_temp_abs_max <- max(ds_temp_abs_max, max(abs(minmax(ds_temp_anomalies[[key]]))))
}
}
# Sequential scale from 0 to max (all warming, no cooling)
anom_min <- min(sapply(ds_temp_anomalies, function(x) minmax(x)[1]))
anom_max <- max(sapply(ds_temp_anomalies, function(x) minmax(x)[2]))
anom_range <- c(anom_min, anom_max)
# Plot 2x2 grid
fname <- paste0(out_dir_graphics, "/dry_season_tmean_change_2x2")
png(paste0(fname, ".png"), width = 12, height = 10, units = "in", res = 300, type = "cairo")
par(mfrow = c(2, 2))
ssp_labels <- list("245" = "SSP2-4.5", "585" = "SSP5-8.5")
for (ssp in ssps) {
for (year in years) {
key <- paste0(ssp, "_", year)
anom <- ds_temp_anomalies[[key]]
mean_val <- round(global(anom, mean, na.rm = TRUE)$mean, 2)
sign_val <- ifelse(mean_val >= 0, "+", "")
plot(anom,
main = paste0(
"Dry season mean temperature change (\u00B0C)\n",
ssp_labels[[ssp]], " ", year, " relative to 1970\u20132000"
),
col = hcl.colors(100, "YlOrRd", rev = TRUE),
range = anom_range,
mar = c(3.5, 3.5, 3.5, 3.5),
xlab = "Longitude (\u00B0E)",
ylab = "Latitude (\u00B0N)",
plg = list(title = "\u00B0C"),
cex.main = 1.0,
cex.lab = 1.0,
box = FALSE
)
lines(adm0, col = "black", lwd = 1.2)
mtext(paste0("National mean: ", sign_val, mean_val, "\u00B0C"),
side = 3, line = -0.5, cex = 0.8, font = 2
)
}
}
dev.off()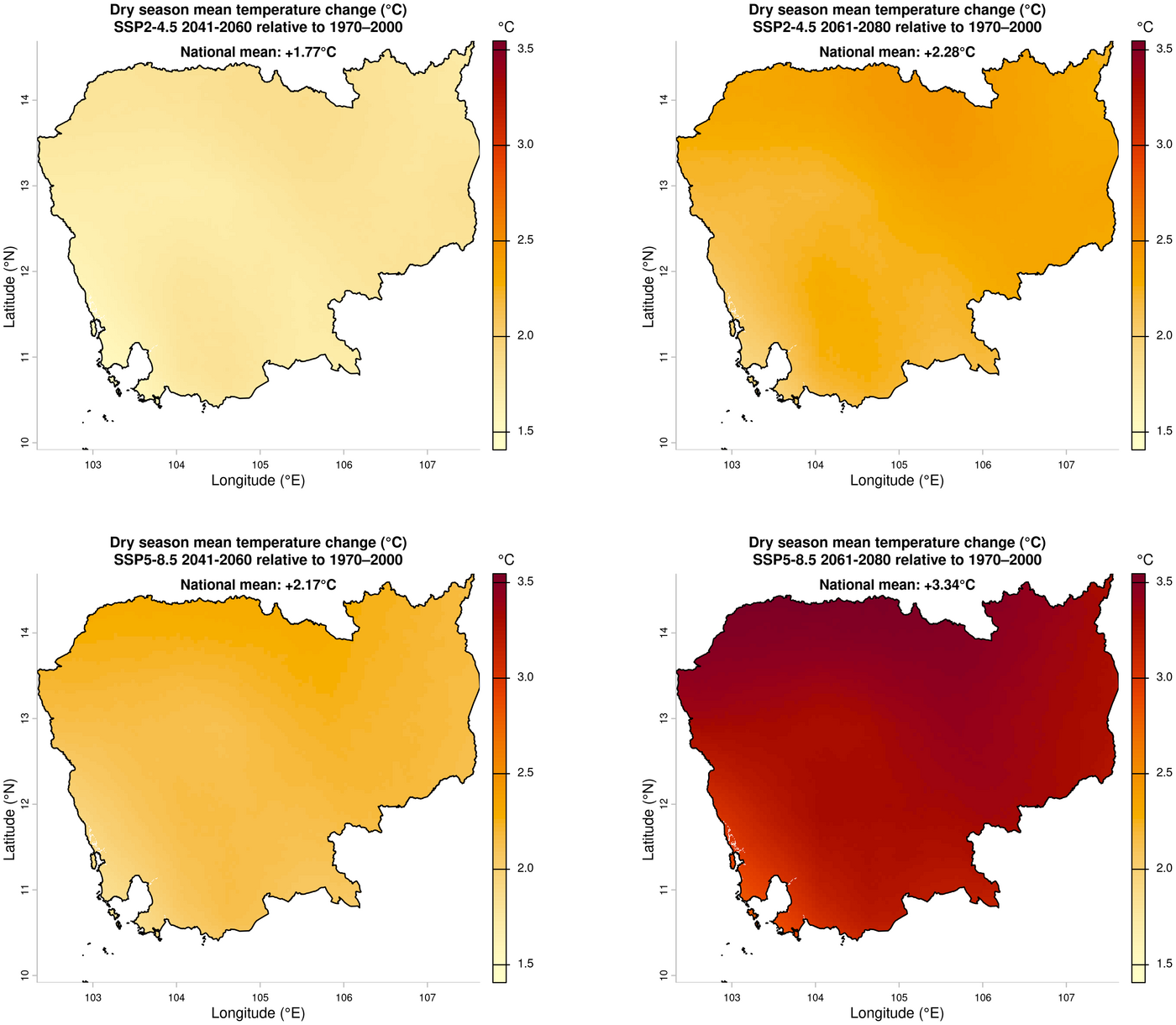
4. Monthly mean precipitation and monsoon periods — historical and future projections
Historical line + SSP2-4.5 and SSP5-8.5 ribbons (GCM spread) in two panels for 2041-2060 and 2061-2080 and shaded wet and dry season monsoon periods. The southwest monsoon is indicated at the start of May and end of October, as noted by (RIMES & UNDP, 2020).
# ============================================================
# Monthly mean precipitation and monsoon periods: Ensemble range plot with monsoon period overlay
# Historical + SSP2-4.5 and SSP5-8.5 ribbons (GCM spread)
# Two panels: 2041-2060 and 2061-2080
# Wet/dry season periods
# SW monsoon onset (May) and cessation (November)
# Two panels: 2041-2060 and 2061-2080
# ============================================================
fname <- paste0(out_dir_graphics, "/cambodia_prec_seasonal_monsoon")
png(paste0(fname, ".png"), width = 12, height = 7, units = "in", res = 300, type = "cairo")
par(mfrow = c(1, 2), mar = c(5, 4.5, 4, 2), mgp = c(2.5, 0.5, 0))
mon <- 1:12
month_labels <- c(
"Jan", "Feb", "Mar", "Apr", "May", "Jun",
"Jul", "Aug", "Sep", "Oct", "Nov", "Dec"
)
ssp_cols <- list("245" = "#EAB139", "585" = "#980002")
ssp_labels <- list("245" = "SSP2-4.5", "585" = "SSP5-8.5")
for (year in years) {
# Future ensemble stats per SSP
ens <- list()
for (ssp in ssps) {
gcm_monthly <- list()
for (g in gcms) {
prec <- rast(paste0(out_dir_future, "/wc2.1_30s_prec_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
prec <- mask(crop(prec, adm0), adm0)
gcm_monthly[[g]] <- as.numeric(global(prec, mean, na.rm = TRUE)$mean)
}
mat <- do.call(rbind, gcm_monthly)
ens[[ssp]] <- list(
mean = colMeans(mat),
min = apply(mat, 2, min),
max = apply(mat, 2, max)
)
}
# Y-axis range
ylim <- range(c(
prec_monthly,
ens[["245"]]$min, ens[["245"]]$max,
ens[["585"]]$min, ens[["585"]]$max
)) + c(-10, 20)
# Empty plot
plot(mon, prec_monthly,
type = "n",
xlab = "",
ylab = "Mean precipitation (mm)",
xaxt = "n",
yaxt = "n",
ylim = ylim,
main = paste0("Monthly mean precipitation and monsoon periods\nCambodia \u2014 historical & future projections (", year, ")"),
bty = "n",
cex.main = 0.9
)
# Wet/dry season background shading
# Wet season: May-Oct
rect(4.5, ylim[1], 10.5, ylim[2],
col = adjustcolor("steelblue", alpha.f = 0.06),
border = NA
)
# Dry season: Jan-Apr
rect(0.5, ylim[1], 4.5, ylim[2],
col = adjustcolor("tan3", alpha.f = 0.06),
border = NA
)
# Dry season: Nov-Dec
rect(10.5, ylim[1], 12.5, ylim[2],
col = adjustcolor("tan3", alpha.f = 0.06),
border = NA
)
# Season labels — midpoints of each season
mtext("Dry", side = 3, at = 2.5, line = -2.5, cex = 0.7, col = "tan4", font = 3)
mtext("Wet", side = 3, at = 7.5, line = -2.5, cex = 0.7, col = "steelblue", font = 3)
mtext("Dry", side = 3, at = 11.5, line = -2.5, cex = 0.7, col = "tan4", font = 3)
# Onset/cessation reference lines
abline(v = 4.5, col = "steelblue", lty = 3, lwd = 1.2)
abline(v = 10.5, col = "steelblue", lty = 3, lwd = 1.2)
# Annotate onset/cessation
text(4.5, ylim[2] - 10, "Onset\n(May)", cex = 0.65, col = "steelblue", adj = 0.5)
text(10.5, ylim[2] - 10, "Cessation\n(Nov)", cex = 0.65, col = "steelblue", adj = 0.5)
axis(1, at = mon, labels = month_labels, cex.axis = 0.85, tcl = -0.2)
axis(2, las = 1, cex.axis = 0.8)
# SSP ribbons
for (ssp in ssps) {
polygon(c(mon, rev(mon)),
c(ens[[ssp]]$max, rev(ens[[ssp]]$min)),
col = adjustcolor(ssp_cols[[ssp]], alpha.f = 0.2),
border = NA
)
}
# Ensemble mean lines
for (ssp in ssps) {
lines(mon, ens[[ssp]]$mean,
col = ssp_cols[[ssp]], lwd = 2, lty = 2
)
}
# Historical line
lines(mon, prec_monthly, col = "black", lwd = 2.5)
# Legend
legend("topleft",
inset = c(0.02, 0.12),
legend = c(
"Historical (1970\u20132000)",
"SSP2-4.5 ensemble mean", "SSP2-4.5 GCM range",
"SSP5-8.5 ensemble mean", "SSP5-8.5 GCM range"
),
col = c(
"black",
"#EAB139", adjustcolor("#EAB139", alpha.f = 0.2),
"#980002", adjustcolor("#980002", alpha.f = 0.2)
),
lwd = c(2.5, 2, 8, 2, 8),
lty = c(1, 2, 1, 2, 1),
bty = "n",
cex = 0.72
)
}
dev.off()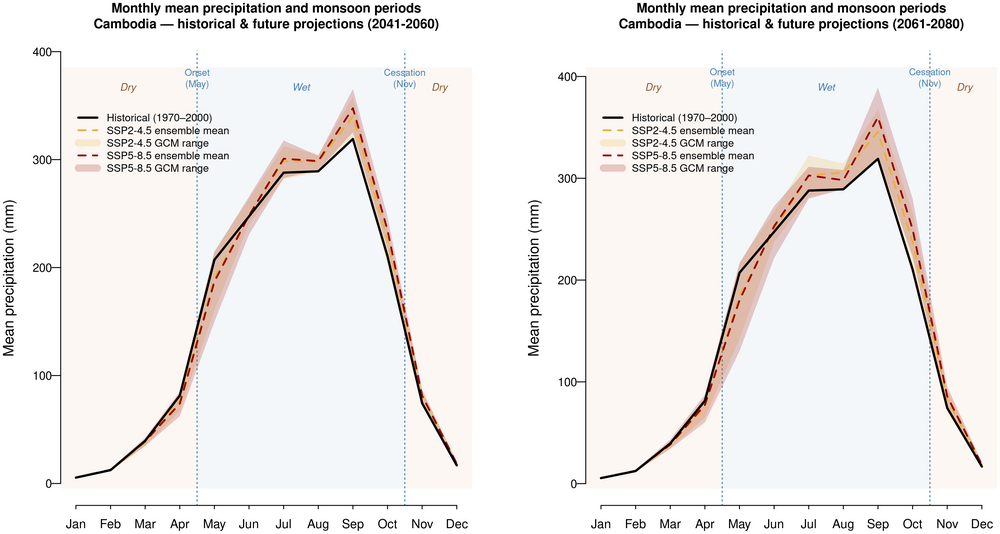
5. Monthly mean Tmax and 35°C threshold — historical and future projections
# ============================================================
# Monthly mean maximum temperature
# Historical and future projections with 35°C threshold line
# Two panels: 2041-2060 and 2061-2080
# ============================================================
# Historical monthly tmax spatial mean
tmax_monthly_hist <- as.numeric(global(hist_tmax_c, mean, na.rm = TRUE)$mean)
fname <- paste0(out_dir_graphics, "/cambodia_monthly_tmax")
png(paste0(fname, ".png"), width = 12, height = 6, units = "in", res = 300, type = "cairo")
par(mfrow = c(1, 2), mar = c(4, 4, 4, 2), mgp = c(2.5, 0.5, 0))
ssp_cols <- list("245" = "#EAB139", "585" = "#980002")
for (year in years) {
# Future ensemble stats for tmax
ens <- list()
for (ssp in ssps) {
gcm_monthly <- list()
for (g in gcms) {
tmax <- rast(paste0(out_dir_future, "/wc2.1_30s_tmax_", g, "_ssp", ssp, "_", year, "_tile-34.tif"))
tmax <- mask(crop(tmax, adm0), adm0)
gcm_monthly[[g]] <- as.numeric(global(tmax, mean, na.rm = TRUE)$mean)
}
mat <- do.call(rbind, gcm_monthly)
ens[[ssp]] <- list(
mean = colMeans(mat),
min = apply(mat, 2, min),
max = apply(mat, 2, max)
)
}
# Y-axis range — ensure 35°C threshold is visible
ylim <- range(c(
tmax_monthly_hist,
ens[["245"]]$min, ens[["245"]]$max,
ens[["585"]]$min, ens[["585"]]$max,
35
)) + c(-0.5, 1)
# Empty plot
plot(mon, tmax_monthly_hist,
type = "n",
xlab = "",
ylab = "Mean maximum temperature (\u00B0C)",
xaxt = "n",
yaxt = "n",
ylim = ylim,
main = paste0("Monthly mean maximum temperature\nCambodia \u2014 historical & future projections (", year, ")"),
bty = "n",
cex.main = 0.9
)
axis(1, at = mon, labels = month_labels, cex.axis = 0.85, tcl = -0.2)
axis(2, las = 1, cex.axis = 0.8)
# 35°C threshold
abline(
h = 35,
col = "firebrick",
lty = 2,
lwd = 1.5
)
text(12, 35,
labels = "35\u00B0C threshold\n\u25B2 months at or above threshold",
col = "firebrick",
cex = 0.7,
adj = c(1, -0.5)
)
# SSP ribbons
for (ssp in ssps) {
polygon(c(mon, rev(mon)),
c(ens[[ssp]]$max, rev(ens[[ssp]]$min)),
col = adjustcolor(ssp_cols[[ssp]], alpha.f = 0.2),
border = NA
)
}
# Ensemble mean lines
for (ssp in ssps) {
lines(mon, ens[[ssp]]$mean,
col = ssp_cols[[ssp]], lwd = 2, lty = 2
)
}
# Historical tmax line
lines(mon, tmax_monthly_hist, col = "black", lwd = 2.5)
# Annotate months exceeding or approaching 35°C
# Indicate months where ensemble max of SSP5-8.5 exceeds 35°C
exceed <- which(ens[["585"]]$max >= 35)
if (length(exceed) > 0) {
points(exceed, rep(35, length(exceed)),
pch = 17, col = "firebrick", cex = 1.0
)
}
# Legend
legend("bottomleft",
inset = c(0.12, 0.02),
legend = c(
"Historical (1970\u20132000)",
"SSP2-4.5 ensemble mean", "SSP2-4.5 GCM range",
"SSP5-8.5 ensemble mean", "SSP5-8.5 GCM range",
"35\u00B0C threshold"
),
col = c(
"black",
"#EAB139", adjustcolor("#EAB139", alpha.f = 0.2),
"#980002", adjustcolor("#980002", alpha.f = 0.2),
"firebrick"
),
lwd = c(2.5, 2, 8, 2, 8, 1.5),
lty = c(1, 2, 1, 2, 1, 2),
bty = "n",
cex = 0.72
)
}
dev.off()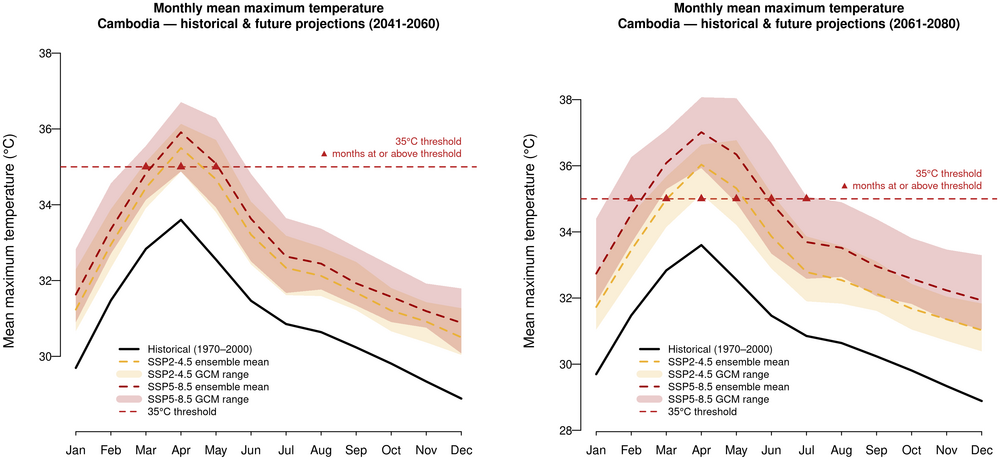
Code
The full Rmd script can be downloaded from GitHub: https://github.com/richardtc/transferable-climate-profile